Synthesis, structure and bioactivity of primary sulfamate-containing natural products.
Mujumdar, P., Bua, S., Supuran, C.T., Peat, T.S., Poulsen, S.A.(2018) Bioorg Med Chem Lett 28: 3009-3013
- PubMed: 29685656
- DOI: https://doi.org/10.1016/j.bmcl.2018.04.038
- Primary Citation of Related Structures:
6C7W, 6C7X - PubMed Abstract:
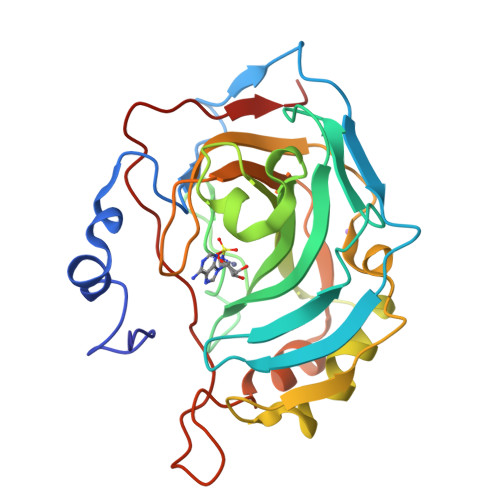
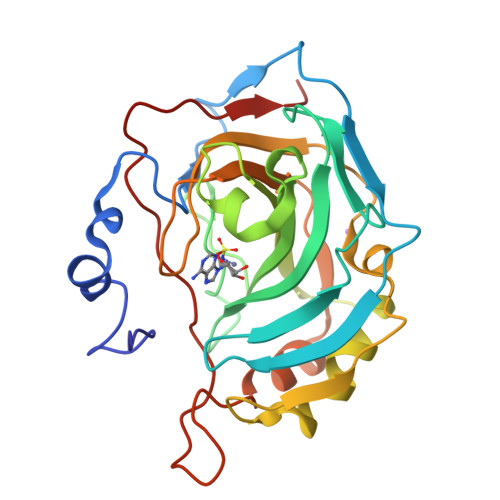
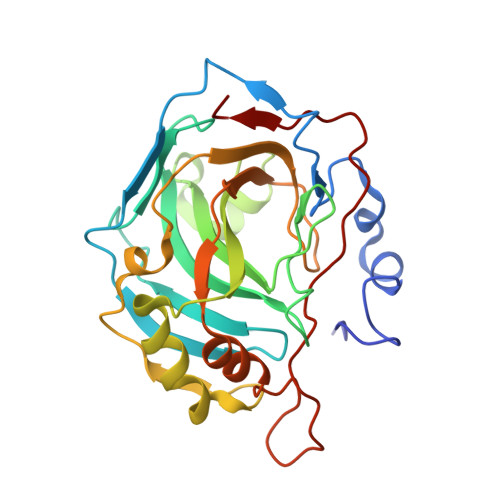
Here we report the synthesis of natural products (NPs) 5'-O-sulfamoyl adenosine 1 and 5'-O-sulfamoyl-2-chloroadenosine 2. As primary sulfamates these compounds represent an uncommon class of NPs, furthermore there are few NPs known that contain a NS bond. Compounds 1 and 2 were evaluated for inhibition of carbonic anhydrases (CA), a metalloenzyme family where the primary sulfamate is known to coordinate to the active site zinc and form key hydrogen bonds with adjacent CA active site residues. Both NPs were good to moderate CA inhibitors, with compound 2 a 20-50-fold stronger CA inhibitor (K i values 65-234 nM) than compound 1. The protein X-ray crystal structures of 1 and 2 in complex with CA II show that it is not the halogen-hydrophobic interactions that give compound 2 a greater binding energy but a slight movement in orientation of the ribose ring that allows better hydrogen bonds to CA residues. Compounds 1 and 2 were further investigated for antimicrobial activity against a panel of microbes relevant to human health, including Gram-negative bacteria (4 strains), Gram-positive bacteria (1 strain) and yeast (2 strains). Antimicrobial activity and selectivity was observed. The minimum inhibitory concentration (MIC) of NP 1 was 10 µM against Gram-positive Staphylococcus aureus and NP 2 was 5 µM against Gram-negative Escherichia coli. This is the first time that NP primary sulfamates have been assessed for inhibition and binding to CAs, with systematic antimicrobial activity studies also reported.
Organizational Affiliation:
Griffith Institute for Drug Discovery, Griffith University, Don Young Road, Nathan, Brisbane, Queensland 4111, Australia.